- こちらでCartogramのことを書いた
- その延長線上で地図も描きたい
- ただ描くのではなくて、座標がほしい
- RのGEOmapパッケージは緯度・経度を指定して地図描図法も指定して地図を表示してくれる関数plotGEOmap()を提供しているのだが、「座標がほしい!」ということで、その関数を以下のように改変(my.retという変数に関する行だけを加えたもの。関数の全部はわからないけれど、座標っぽいところを変数に入れて返り値にしてみたらうまく行った)
my.plotGEOmapXY <-
function (MAP, LIM = c(-180, -90, 180, 90), PROJ = list(), PMAT = NULL,
add = TRUE, SEL = NULL, GRID = NULL, GRIDcol = 1, MAPcol = NULL,
MAPstyle = NULL, border = NA, cenlon = 0, shiftlon = 0, linelty = 1,
linelwd = 1, ptpch = ".", ptcex = 1, NUMB = FALSE, ...)
{
if (missing(cenlon)) {
cenlon = 0
}
if (missing(GRID)) {
GRID = NULL
}
if (missing(GRIDcol)) {
GRIDcol = 1
}
if (missing(PMAT)) {
PMAT = NULL
}
if (missing(SEL)) {
SEL = NULL
}
if (missing(linelty)) {
linelty = 1
}
if (missing(NUMB)) {
NUMB = FALSE
}
if (missing(linelwd)) {
linelwd = 1
}
if (missing(ptpch)) {
ptpch = "."
}
if (missing(ptcex)) {
ptcex = 1
}
if (missing(MAPcol)) {
MAPcol = NULL
}
if (missing(MAPstyle)) {
MAPstyle = NULL
}
if (missing(border)) {
border = NA
}
if (missing(shiftlon)) {
shiftlon = 0
}
pctexp = 0.01
if (missing(LIM)) {
lon = RPMG::fmod(MAP$POINTS$lon - shiftlon, 360)
RLON = expandbound(range(RPMG::fmod(MAP$POINTS$lon, 360)),
pctexp)
RLAT = expandbound(range(MAP$POINTS$lat), pctexp)
LIM = c(RLON[1], RLAT[1], RLON[2], RLAT[2])
}
else {
if (is.null(LIM)) {
RLON = expandbound(range(RPMG::fmod(MAP$POINTS$lon,
360)), pctexp)
RLAT = expandbound(range(MAP$POINTS$lat), pctexp)
LIM = c(RLON[1], RLAT[1], RLON[2], RLAT[2])
}
if (is.list(LIM)) {
lon = RPMG::fmod(LIM$lon - shiftlon, 360)
lat = LIM$lat
LIM = c(min(RPMG::fmod(lon, 360)), min(lat), max(fmod(lon,
360)), max(lat))
}
}
LLlim = list(lat = LIM[c(2, 4)], lon = LIM[c(1, 3)])
if (missing(PROJ)) {
MAPCENLAT = mean(LLlim$lat)
MAPCENLON = median(LLlim$lon)
PROJ = setPROJ(type = 2, LAT0 = MAPCENLAT, LON0 = MAPCENLON,
LATS = NULL, LONS = NULL, DLAT = NULL, DLON = NULL,
FN = 0)
}
if (missing(add)) {
add = FALSE
}
if (is.null(MAP$POINTS$z)) {
MAP$POINTS$z = rep(0, length(MAP$POINTS$lat))
}
if (is.null(MAP$POINTS$x)) {
MAPXY = GLOB.XY(MAP$POINTS$lat, RPMG::fmod(MAP$POINTS$lon -
shiftlon, 360), PROJ)
MAXDISTX = max(MAPXY$x) - min(MAPXY$x)
if (is.null(PMAT)) {
MAP$POINTS$x = MAPXY$x
MAP$POINTS$y = MAPXY$y
}
else {
tem = trans3d(MAPXY$x, MAPXY$y, MAP$POINTS$z, PMAT)
MAP$POINTS$x = tem$x
MAP$POINTS$y = tem$y
}
STRKXYLL = GLOB.XY(MAP$STROKES$LAT1, RPMG::fmod(MAP$STROKES$LON1 -
shiftlon, 360), PROJ)
STRKXYUR = GLOB.XY(MAP$STROKES$LAT2, RPMG::fmod(MAP$STROKES$LON2 -
shiftlon, 360), PROJ)
STRKXYUL = GLOB.XY(MAP$STROKES$LAT2, RPMG::fmod(MAP$STROKES$LON1 -
shiftlon, 360), PROJ)
STRKXYLR = GLOB.XY(MAP$STROKES$LAT1, RPMG::fmod(MAP$STROKES$LON2 -
shiftlon, 360), PROJ)
if (is.null(PMAT)) {
MAP$STROKES$x1 = STRKXYLL$x
MAP$STROKES$y1 = STRKXYLL$y
}
else {
tem = trans3d(STRKXYLL$x, STRKXYLL$y, rep(0, length(STRKXYLL$y)),
PMAT)
MAP$STROKES$x1 = tem$x
MAP$STROKES$y1 = tem$y
}
if (is.null(PMAT)) {
MAP$STROKES$x2 = STRKXYUR$x
MAP$STROKES$y2 = STRKXYUR$y
}
else {
tem = trans3d(STRKXYUR$x, STRKXYUR$y, rep(0, length(STRKXYUR$y)),
PMAT)
MAP$STROKES$x2 = tem$x
MAP$STROKES$y2 = tem$y
}
}
XYLIM = GLOB.XY(LLlim$lat, LLlim$lon, PROJ)
LLlim$x = XYLIM$x
LLlim$y = XYLIM$y
xrange = abs(diff(range(LLlim$x, na.rm = TRUE)))
yrange = abs(diff(range(LLlim$y, na.rm = TRUE)))
Kstroke = length(MAP$STROKES$num)
FORCE = FALSE
if (!is.null(SEL)) {
if (length(SEL) < 0) {
FORCE = TRUE
SEL = 1:length(MAP$STROKES$num)
}
}
if (FORCE == FALSE) {
IN = KINOUT(MAP, LLlim, projtype = 2)
}
else {
IN = 1:length(MAP$STROKES$num)
}
if (!is.null(SEL)) {
slin = which(IN %in% SEL)
if (length(slin) >= 1) {
IN = IN[slin]
}
}
minx = Inf
maxx = -Inf
miny = Inf
maxy = -Inf
if (length(IN) < 1) {
print("No map strokes in target")
return(0)
}
for (i in IN) {
j1 = MAP$STROKES$index[i] + 1
j2 = j1 + MAP$STROKES$num[i] - 1
if (j1 > 0 & j2 > 0 & j2 - j1 >= 0) {
JEC = j1:j2
x = MAP$POINTS$x[JEC]
y = MAP$POINTS$y[JEC]
x[x < LLlim$x[1] | x > LLlim$x[2]] = NA
y[y < LLlim$y[1] | y > LLlim$y[2]] = NA
minx = min(c(minx, x), na.rm = TRUE)
maxx = max(c(maxx, x), na.rm = TRUE)
miny = min(c(miny, y), na.rm = TRUE)
maxy = max(c(maxy, y), na.rm = TRUE)
}
else {
next
}
}
if (!is.null(MAPcol)) {
MAP$STROKES$col = rep(MAPcol, length = length(MAP$STROKES$col))
}
if (!is.null(MAPstyle)) {
MAP$STROKES$style = rep(MAPstyle, length = length(MAP$STROKES$style))
}
if (add == FALSE) {
plot(range(LLlim$x), range(LLlim$y), asp = 1, type = "n",
...)
}
if (!is.null(GRID)) {
addLLXY(GRID$lats, GRID$lons, PMAT = PMAT, GRIDcol = GRIDcol,
LABS = 0, BORDER = 0, PROJ = PROJ)
}
my.ret <- matrix(0,0,2)
for (i in IN) {
j1 = MAP$STROKES$index[i] + 1
j2 = j1 + MAP$STROKES$num[i] - 1
if ((j1 > 0 & j2 > 0 & j2 - j1 >= 0)) {
JEC = j1:j2
}
else {
next
}
if (NUMB == TRUE) {
points(MAP$POINTS$x[j1], MAP$POINTS$y[j1])
my.ret <- rbind(my.ret,cbind(MAP$POINTS$x[j1], MAP$POINTS$y[j1]))
text(MAP$POINTS$x[j1], MAP$POINTS$y[j1], labels = i,
pos = 3)
}
if (MAP$STROKES$style[i] == 1) {
points(MAP$POINTS$x[JEC], MAP$POINTS$y[JEC], col = MAP$STROKES$col[i],
pch = ptpch, cex = ptcex)
my.ret <- rbind(my.ret,cbind(MAP$POINTS$x[JEC], MAP$POINTS$y[JEC]))
}
if (MAP$STROKES$style[i] == 2) {
x = MAP$POINTS$x[JEC]
y = MAP$POINTS$y[JEC]
my.ret <- rbind(my.ret,cbind(x,y))
xd = abs(diff(c(x, x[1])))
wwx = which(xd > 0.9 * xrange)
yd = abs(diff(c(y, y[1])))
wwy = which(yd > 0.8 * yrange)
if ((!is.null(wwx) & length(wwx) > 0) | (!is.null(wwy) &
length(wwy) > 0)) {
if (length(wwx) > 0) {
zy = insertNA(y, wwx)
zx = insertNA(x, wwx)
lines(zx, zy, col = MAP$STROKES$col[i], lty = linelty,
lwd = linelwd)
}
if (length(wwy) > 0) {
zy = insertNA(y, wwy)
zx = insertNA(x, wwy)
lines(zx, zy, col = MAP$STROKES$col[i], lty = linelty,
lwd = linelwd)
}
}
else {
x[MAP$POINTS$x[JEC] < LLlim$x[1] | MAP$POINTS$x[JEC] >
LLlim$x[2]] = NA
y[MAP$POINTS$y[JEC] < LLlim$y[1] | MAP$POINTS$y[JEC] >
LLlim$y[2]] = NA
lines(x, y, col = MAP$STROKES$col[i], lty = linelty,
lwd = linelwd)
}
}
if (MAP$STROKES$style[i] == 3) {
x = MAP$POINTS$x[JEC]
y = MAP$POINTS$y[JEC]
x = c(x, x[1])
y = c(y, y[1])
x[MAP$POINTS$lon[JEC] < LLlim$lon[1] | MAP$POINTS$lon[JEC] >
LLlim$lon[2]] = NA
y[MAP$POINTS$lat[JEC] < LLlim$lat[1] | MAP$POINTS$lat[JEC] >
LLlim$lat[2]] = NA
polygon(x, y, border = border, col = MAP$STROKES$col[i])
my.ret <- rbind(my.ret,cbind(x,y))
}
}
invisible(IN)
return(my.ret)
}
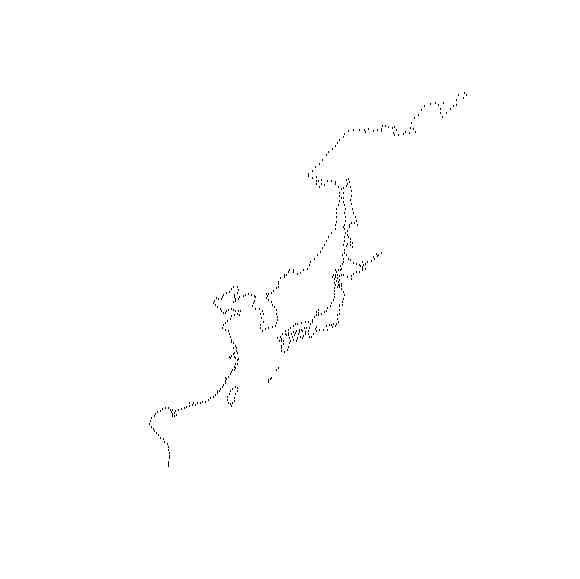
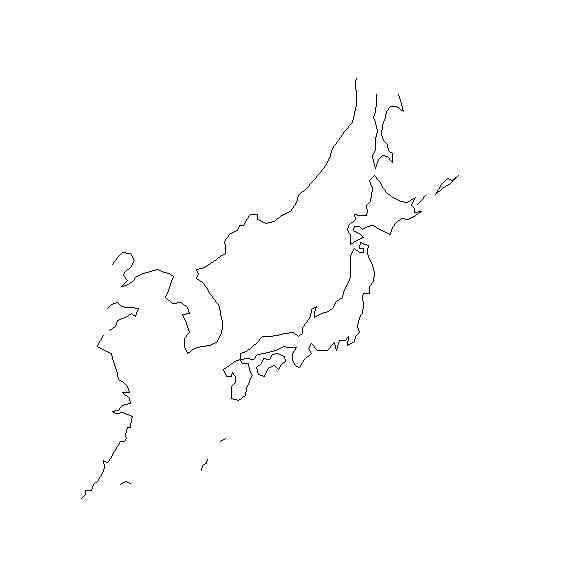
library(geomapdata)
data(worldmap)
KAMlat = c(25, 50)
KAMlon = c(120, 150)
PLOC=list(LON=KAMlon,LAT=KAMlat)
PLON = seq(from=KAMlon[1], to=KAMlon[2], by=2)
PLAT = seq(from=KAMlat[1], to=KAMlat[2], by=2)
proj = setPROJ(1, LON0=mean(KAMlon), LAT0=mean(KAMlat))
xy = GLOB.XY(KAMlat, KAMlon , proj)
kbox=list(x=range(xy$x, na.rm=TRUE), y=range(xy$y, na.rm=TRUE))
plot(kbox$x,kbox$y, type='n', axes=FALSE, xlab="", ylab="", asp=1)
mmm <- my.plotGEOmapXY(worldmap, LIM=c(KAMlon[1], KAMlat[1], KAMlon[2],KAMlat[2]), add=TRUE, PROJ=proj, axes=FALSE, xlab="", ylab="" )
dev.new()
plot(mmm,cex=0.01,axes=FALSE, xlab="", ylab="", asp=1)
- 地図座標が得られたので、標本の出身地を地図上座標として取り、また、標本の属性からMDSを経由して属性に関する座標をとり、それに基づいて地図を歪めて見る
- データがよいものではないのでむちゃくちゃだが、処理を残しておく、ということで
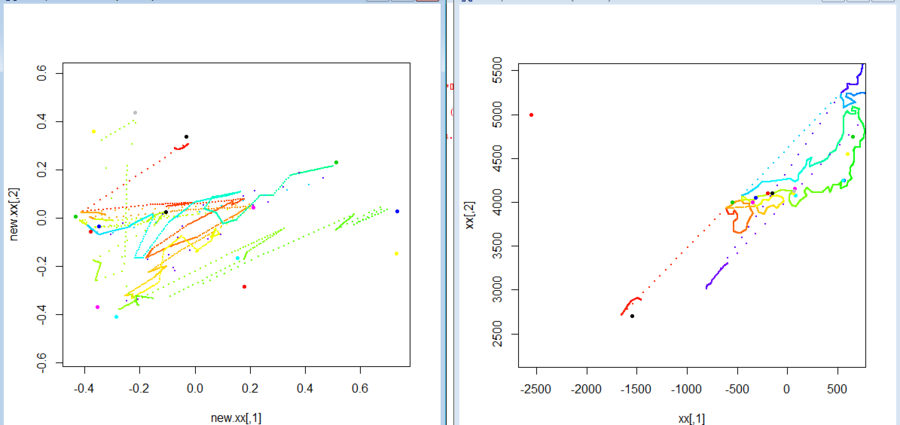
> seyana.data.1
奈良 高槻 大阪 吹田
1 0.3 0.70 0.8 0.8
2 0.7 0.70 0.5 0.2
3 0.5 0.20 0.8 0.3
4 0.9 0.90 0.7 0.7
5 1.0 0.98 0.4 0.5
6 1.0 1.00 0.4 0.6
7 0.9 0.70 1.0 0.9
8 0.8 0.50 0.9 1.0
9 0.7 0.70 0.6 0.7
10 0.8 0.90 0.5 0.9
11 0.9 0.99 0.8 0.7
12 0.4 0.10 0.5 0.2
13 0.6 0.50 0.2 0.7
14 0.8 0.40 0.7 0.4
15 0.5 0.10 0.3 0.2
> seyana.coords
V1 V2
1 -1550 2700
2 -2550 5000
3 650 4750
4 -320 4050
5 80 4080
6 70 4150
7 60 4100
8 65 4125
9 -150 4100
10 -200 4100
11 -550 4000
12 560 4250
13 550 4250
14 -350 4000
15 600 4550
library(geomapdata)
data(worldmap)
KAMlat = c(25, 50)
KAMlon = c(120, 150)
PLOC=list(LON=KAMlon,LAT=KAMlat)
PLON = seq(from=KAMlon[1], to=KAMlon[2], by=2)
PLAT = seq(from=KAMlat[1], to=KAMlat[2], by=2)
proj = setPROJ(1, LON0=mean(KAMlon), LAT0=mean(KAMlat))
xy = GLOB.XY(KAMlat, KAMlon , proj)
kbox=list(x=range(xy$x, na.rm=TRUE), y=range(xy$y, na.rm=TRUE))
plot(kbox$x,kbox$y, type='n', axes=FALSE, xlab="", ylab="", asp=1)
mmm <- my.plotGEOmapXY(worldmap, LIM=c(KAMlon[1], KAMlat[1], KAMlon[2],KAMlat[2]), add=TRUE, PROJ=proj, axes=FALSE, xlab="", ylab="" )
dev.new()
plot(mmm,cex=0.01,axes=FALSE, xlab="", ylab="", asp=1)
K <- 20
mmm.2 <- matrix(0,0,2)
for(i in 300:length(mmm[,1])){
mmm.2 <- rbind(mmm.2,cbind(seq(from=mmm[i-1,1],to=mmm[i,1],length=K),seq(from=mmm[i-1,2],to=mmm[i,2],length=K)))
}
X <- as.matrix(seyana.coords)
library(geometry)
delaunay.X <- delaunayn(X)
plot(X)
delaunay.X.2 <- rbind(delaunay.X[,1:2],delaunay.X[,2:3],delaunay.X[,c(3,1)])
segments(X[delaunay.X.2[,1],1],X[delaunay.X.2[,1],2],X[delaunay.X.2[,2],1],X[delaunay.X.2[,2],2])
D <- as.matrix(dist(seyana.data.1))
X.. <- cmdscale(D)
plot(X..)
xx <- mmm.2
judge.inside.2 <- function(x,V){
n <- length(x)
K <- matrix(V[1:n,],ncol=n)
K. <- t(K)-V[n+1,]
tmp <- solve(K.,x-V[n+1,])
c(tmp,1-sum(tmp))
}
new.xx <- matrix(NA,length(xx[,1]),length(xx[1,]))
for(i in 1:length(delaunay.X[,1])){
tmp.v <- delaunay.X[i,]
V <- X[tmp.v,]
tmp <- t(apply(xx,1,judge.inside.2,V))
tmp.more.0 <- apply(tmp>=0,1,all)
insiders <- which(tmp.more.0)
if(length(insiders) > 0){
new.xx[insiders,] <- tmp[insiders,] %*% X..[tmp.v,]
}
}
name.city <- c("台中","中国","盛岡","因島","奈良","高槻","大阪","吹田","善通寺","福山","福岡","青山","洗足","松山","宮城")
xlim <- ylim <- range(c(xx))
col <- rainbow(length(mmm.2[,1]))
x.diff <- max(X[,1])-min(X[,1])
y.diff <- max(X[,2])-min(X[,2])
x.mean <- mean(range(X[,1]))
y.mean <- mean(range(X[,2]))
xlim <- c(-0.5,0.5)*max(x.diff,y.diff)+x.mean
ylim <- c(-0.5,0.5)*max(x.diff,y.diff)+y.mean
plot(xx,pch=20,cex=0.01,col=col,xlim=xlim,ylim=ylim)
points(X,pch=20,cex=1,col=1:length(X[,1]))
dev.new()
x.diff <- max(X..[,1])-min(X..[,1])
y.diff <- max(X..[,2])-min(X..[,2])
x.mean <- mean(range(X..[,1]))
y.mean <- mean(range(X..[,2]))
xlim <- c(-0.5,0.5)*max(x.diff,y.diff)+x.mean
ylim <- c(-0.5,0.5)*max(x.diff,y.diff)+y.mean
plot(new.xx,pch=20,cex=0.01,col=col,xlim=xlim,ylim=ylim)
points(X..,pch=20,cex=1,col=1:length(X..[,1]))


